Phylogenetic analysis
The amino acid sequence of GeCas12a2 was aligned with other Cas12a2 nuclease sequences15,59 using Clustal Omega60. The alignment was trimmed with ClipKIT61 and used to reconstruct a phylogeny with IQ-TREE v2.0.3 (-m MFP -T 8 -B 1000)62, using a maximum-likelihood approach. The phylogeny included Cas12a orthologues and used Cas12c as an outgroup. Branch confidence was reported as ultrafast bootstrap values (ranging from 0 to 100).
PFS library preparation
A PFS-containing plasmid library (CBS-6873) was constructed to include a target sequence (Supplementary Tables 8 and 9) followed by five randomized nucleotides (NNNNN). The target sequence was placed under a PJ23119 promoter and cloned into a low-copy sc101 origin plasmid (about five copies per cell). The library was generated by amplifying a target-encoding plasmid with primers ODpr23 and ODpr24 (Supplementary Tables 9 and 10). The forward primer contained a 5-nt randomized overhang. The PCR product was treated with DpnI to remove template DNA, ligated, and electroporated into Escherichia coli (E. coli) TOP10, yielding over two million transformants (approximately 2,000-fold library coverage).
PFS library depletion
The PFS preference of the GeCas12a2 nuclease (CBS-6874) was assessed by targeting the CBS-6873 PFS plasmid library with a CAO1-targeting CRISPR RNA (crRNA) plasmid (CBS-6875) or a non-targeting crRNA plasmid (CBS-6876). The GeCas12a2 nucleotide sequence was codon-optimized for expression in E. coli. The nuclease-encoding sequence was under a T7 promoter, whereas crRNA was expressed from PJ23119. Escherichia coli BL21(AI) cells containing nuclease and crRNA plasmids were electroporated with the PFS plasmid library. Each electroporation used approximately 500 ng of library plasmid DNA in 50 µl competent cells, followed by recovery in lysogeny broth medium containing 0.1 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) and 0.2% l-arabinose. Overnight cultures (of approximately two million transformants) ensured greater-than-2,000-fold library coverage. Plasmids were purified with ZymoPURE II Plasmid Midiprep Kit (D4201).
PFS library sequencing and analysis
Purified plasmids from targeting and non-targeting conditions were PCR-amplified using primers ODpr55 and ODpr56 (Supplementary Table 10) with KAPA HIFI HotStart polymerase (KK2601) for 20 cycles at 64.5 °C, following the manufacturer’s protocol. Indexed PCR products were sequenced on Illumina NovaSeq 6000 (paired-end, 150 bp reads), at least two million reads each. Raw FASTQ files were processed with Trimmomatic v.0.39 (parameters: ILLUMINACLIP:TruSeq3-PE.fa:2:30:10, LEADING:3, TRAILING:3, SLIDINGWINDOW:4:15). Paired-end reads were merged with BBMerge (qtrim = t trimq = 10 minlength = 20). Sequences containing motifs matching TTCCTTCAGGTGTTGCTCCA (…..)GGTGAGTTCT were extracted, excluding sequences with N bases or Phred scores of below 20. Depletion scores were calculated using the following formula: depletion = sum(non-target)/sum(target) × count(target)/count(non-target). log2 fold-change values for depletion scores were computed for nucleotides at PFS positions (−1 to −5). Scatterplots visualizing PFS preferences were generated using Matplotlib in Python.
Coexpression of gRNA and FLAG-tagged GeCas12a2 or FLAG-tagged FnCas12a
Electrocompetent E. coli BW25113 cells were transformed with a GeCas12a2-FLAG expression plasmid and a plasmid containing the gRNA’s sequence (Supplementary Table 9). Newly transformed cells were grown on lysogeny broth agar plates containing 25 µg ml–1 kanamycin (Kan) and 100 µg ml–1 ampicillin (Amp) for 12–16 h at 37 °C. A single colony was selected and used to start an overnight culture, from which a 120 ml culture of lysogeny broth–Amp–Kan medium was inoculated to a starting OD600 of 0.05, and then incubated at 37 °C at 200 r.p.m. When OD600 = 0.2 was reached, the cultivation temperature was lowered from 37 °C to 21 °C. Once the growth reached an OD600 of 0.5, expression was induced by adding L-arabinose (to a final concentration of 0.2%). Cells were collected by centrifugation after 16–18 h at 21 °C.
Purification of FLAG-tagged GeCas12a2 or FnCas12a–gRNA RNP complex
Escherichia coli cell pellets (90 optical density volumes) were resuspended in 3 ml Tris-buffered saline (TBS) (150 mM NaCl, 50 mM Tris/HCl pH7.4) containing a mix of protease inhibitors (Complete, ethylenediaminetetraacetic acid-free; Sigma Aldrich), and then lysed by sonication on ice (Branson Sonifier 250 with four ultrasonic cycles, 50% duty cycle, 2.5 output for 30 s). The lysate was cleared by centrifugation (5,525 × g for 10 min at 4 °C) and filtered through a 0.22 µm polyethersulfone membrane (Merck Chemicals). The cleared lysate was divided into two 1.5 ml aliquots and added to 120 µl of magnetic anti-FLAG beads (Pierce Anti-DYKDDDDK magnetic agarose; Thermo Fisher Scientific) that were pre-washed twice with TBS buffer. The lysate was incubated with the magnetic anti-FLAG beads overnight on a rotating wheel at 4 °C, allowing the FLAG-tagged RNP complexes to bind to the magnetic anti-FLAG beads. The magnetic beads were then washed three times with ice-cold TBS buffer and once with double-distilled water (with the magnetic beads fixed to a magnetic rack in between each wash). The supernatant was completely removed, and the FLAG-tagged RNPs were eluted using 1.5 mg ml–1 of a triple FLAG peptide (Pierce 3×DYKDDDDK Peptide; Thermo Fisher Scientific) resuspended in 50 µl phosphate-buffered saline (PBS) (Dulbecco’s PBS without Ca2+ and Mg2+; PAN Biotech) on a slowly shaking platform for 30 min at room temperature. The purified FLAG-tagged RNPs were collected by fixing the magnetic beads in a magnetic rack and pipetting out the supernatant. The purified RNPs were quantified by measuring the absorbance at 280 nm (DeNovix reader), before being stored at 4 °C.
Expression of apo and catalytically dead GeCas12a2 and SuCas12a2
Apo GeCas12a2 and apo SuCas12a2 were expressed similarly to the expression method of SuCas12a2 described in a past work16. In brief, the gecas12a2 gene sequence was cloned into a 2S-T pET plasmid in frame with an N-terminal HIS-SUMO tag. SuCas12a2 was cloned into a pACYC plasmid with a N-term-hexa HIS tag. Both plasmids were separately transformed into chemically competent E. coli HMS-174(DE3) cells. Protein expression for GeCas12a2 and SuCas12a2 was performed using similar methods. A single colony was used to inoculate 60 ml of lysogeny broth medium for an overnight (16–18 h) growth at 37 °C with shaking at 200 r.p.m. The starter growth (20 ml) was then used to inoculate 1 l lysogeny broth medium containing 100 µg ml–1 ampicillin. The 1 l cultures were grown to an OD600 of 0.5–0.6 at 37 °C, and then cold shocked on ice for 20 min before being induced with 0.1 mM IPTG; this was followed by growing the cells for 16–18 h at 18 °C. Cell pellets were collected by centrifugation and stored at −80 °C.
Purification of apo and catalytically dead GeCas12a2 and SuCas12a2
Apo GeCas12a2/Ge(d)Cas12a2 and apo SuCas12a2 were purified similarly to the purification method of SuCas12a2 outlined in a previous work16. In brief, collected cell pellets were thawed on ice for 20 min before being resuspended in 50 ml of lysis buffer (25 mM Tris pH 7.2, 500 mM NaCl, 2 mM MgCl2, 10 mM imidazole, 10% glycerol) containing protease inhibitors (2 μg ml−1 aprotinin, 10 μM leupeptin, 1.0 μg ml−1 pepstatin) and 1 mg ml−1 lysozyme; they were then incubated for 20 min on ice with gentle rocking. Cells were further lysed by sonication (at 3/50 for 25 min) and clarified by centrifugation at 36,400 g for 35 min. Clarified lysate was added to 5 ml of Ni-NTA resin and batch bound at 4 °C for 30 min, followed by running the lysate over the resin for a second time. The resin was washed with 500 ml of Ni-NTA wash buffer (25 mM Tris pH 7.2, 2 M NaCl, 2 mM MgCl2, 10 mM imidazole, 10% glycerol) and eluted with nickel elution buffer (25 mM Tris pH 7.2, 500 mM NaCl, 2 mM MgCl2, 250 mM imidazole, 10% glycerol). Using a HiPrep 26/10 desalting column (Cytiva), elutions from the nickel column were desalted into low-salt buffer (25 mM Tris pH 7.2, 50 mM NaCl, 2 mM MgCl2, 10% glycerol). GeCas12a2’s low-salt buffer contained 200 mM, rather than 50 mM, NaCl to help with solubility. Furthermore, GeCas12a2 was treated for tag cleavage by adding 1:50 tobacco etch virus (TEV) protease and dithiothreitol (DTT) to GeCas12a2 for 2 h at room temperature. Both apo GeCas12a2 and apo SuCas12a2 were applied to a Hitrap SP HP cation-exchange column and eluted with a gradient of high-salt buffer (25 mM Tris pH 7.2, 1.0 M NaCl, 2 mM MgCl2, 10% glycerol). Fractions containing apo GeCas12a2 or apo SuCas12a2 were concentrated to 1 ml using a 100 kDa MWKO concentrator. Concentrated protein was loaded over a HiLoad 26/600 Superdex 200 pg column using size-exclusion chromatography buffer (100 mM HEPES pH 7.2, 150 mM KCl, 2 mM MgCl2, 10% glycerol). Peak fractions containing apo GeCas12a2 or apo SuCas12a2 were concentrated to a working concentration again using a 100 kDa MWKO concentrator before aliquoting and flash freezing with liquid nitrogen and stored at −80 °C.
FAM-labelled collateral cleavage assays
Collateral cleavage assays were performed as described previously15. In brief, 20 µl reactions containing 1× NEB 3.1 buffer (100 mM NaCl, 50 mM Tris-HCl, 10 mM MgCl2, 100 µg ml–1 bis(trimethylsilyl)acetamide (BSA), pH 7.9) were made by combining 300 nM crRNA with 250 nM of GeCas12a2 or SuCas12a2, along with 100 nM of 5′-FAM labelled collateral substrate (ssDNA, dsDNA or ssRNA). Finally, 250 nM of target RNA was added to initiate the reaction and incubated at 37 °C for 1 h. Reactions were quenched with phenol, and nucleic acid was purified by phenol–chloroform extraction. The cleaved nucleic acid was analysed by 12% urea-polyacrylamide gel electrophoresis (urea-PAGE) and visualized for fluorescein fluorescence. Guide and target RNA were in vitro transcribed to determine dsDNA cleavage when targeting mRNA transcripts (GAPDH, EGFP, MALAT1, HPV18.6, HPV18.7).
Yeast cell culture and quantification
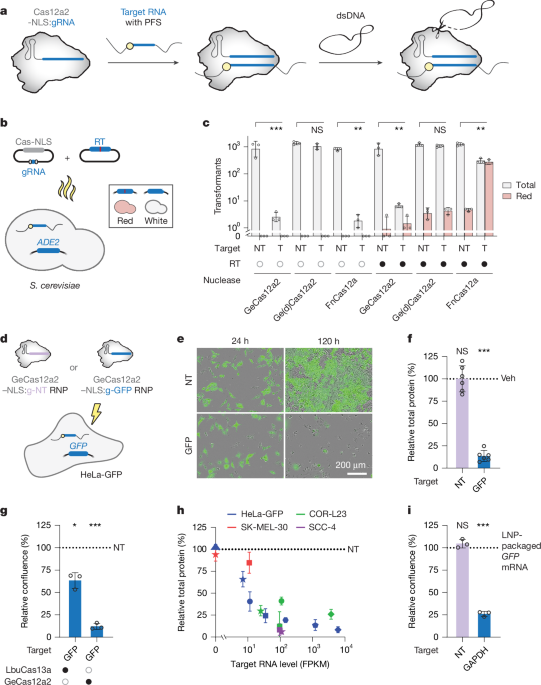
Chemically competent S. cerevisiae S288c cells were transformed using the high-efficiency lithium acetate/single-stranded carrier DNA/PEG method63, using 1 µg of the nuclease/gRNA expression plasmid and 500 ng HDR-template (Supplementary Tables 11 and 12). Following regeneration, transformed cells were plated on selective yeast extract peptone dextrose agar plates containing 50 µg ml–1 geneticin (G418) and incubated for at least two days at 30 °C. The colonies were then counted and grouped according to their colour, with a successful knockout being determined by the red colour of the clone.
Mammalian cell culture
HeLa, HeLa-GFP, HeLa-RFP, HEK293T, HEK293-GFP, SiHa and U2OS cells were cultured in Dulbecco’s modification of Eagle’s medium (DMEM) supplemented with 10% serum, glucose, L-glutamine and sodium pyruvate. HeLa-GFP and HeLa-RFP cultures were supplemented with 10 µg ml–1 blasticidin (InvivoGen) and 2.5 µg ml–1 puromycin (InvivoGen), respectively, to maintain reporter selection. COR-L23 cells were cultured in RPMI 1640 medium supplemented with 10% serum and l-glutamine. SCC-4 cells were cultured in Ham’s F12/DMEM (1:1) containing 20% serum, L-glutamine and 50 µg ml–1 hydrocortisone (Sigma-Aldrich). SK-MEL-30 were cultured in DMEM supplemented with 10% serum, glucose and 1× GlutaMAX (Fisher Scientific). PC3-GFP cells were cultured in F12K medium (Kaighn’s modification) supplemented with 10% serum and L-glutamine. NCI-H23 cells were cultured in RPMI 1640 (ATCC Modification) supplemented with 10% serum, glucose, L-glutamine, sodium pyruvate and sodium bicarbonate. All media were further supplemented with broad spectrum antimicrobials and tested monthly for detection of mycoplasma. All cultures were incubated at 37 °C with 5% CO2 (refer to Supplementary Table 8 for a complete list of cell lines and origins).
Electroporation of CRISPR RNPs to mammalian cells
Cells (1 × 105) were transfected with RNPs (GeCas12a2, SuCas12a2, LbuCas13a, FnCas12a) made by combining a 1:1 molar ratio of 50 pmol of nuclease and gRNA at room temperature for 30 min. For 5 × 105 and 1 × 106 cell reactions, 250 pmol of nuclease and gRNA were incubated together. The RNP complexes were transfected using the Neon and Neon NxT electroporation systems (Thermo Fisher Scientific). Device settings were: 1,300 V/30 ms/1 pulse for HeLa cell types; 1,200 V/40 ms/1 pulse for HEK293 cell types; 1,400 V/15 ms/2 pulses for U2OS cell types; 1,250 V/30 ms/1 pulse for PC3 cell types; and 1,400 V/30 ms/1 pulse for H23 cell types. COR-L23, SCC-4 and SK-MEL-30 cells were electroporated using the NEPA21 electroporator (Nepa Gene) with the following conditions: 150 V for 5 ms, 175 V for 5 ms and 150 V for 7.5 ms, respectively. After pulsing the cells, samples were plated according to assay type or further cultured for downstream treatment and analysis.
Imaging, cell counting and intensity quantification by Incucyte tracking
Following payload delivery, cells were seeded in 96-well or 24-well plates and live-imaged every 4 h for up to five days in phase contrast and, when relevant, GFP channels (300 ms acquisition) using the Incucyte SX5 system (Sartorius) and Incucyte Base Analysis Software (v2023A). Four to sixteen images were acquired per well, from which representative figures were prepared. Incucyte software-based cell quantifications include the phase contrast confluence, phase contrast object count per image, GFP object count per image and the total integrated GFP intensity per image (refer to Figs. 1e,g,i, 5e,g,i,j and Extended Data Fig. 4d,e).
Total cellular protein quantification by SRB
Cells in 96-well plates were fixed with 10% trichloroacetic acid for 30 min on ice and stained with 0.4% sulforhodamine B (Sigma Aldrich) in 1% acetic acid solution for 10 min at room temperature. After washing several times with 1% acetic acid, the 96-well plates were allowed to dry at room temperature. Sulforhodamine B dye was extracted by incubation in chilled 10 mM TRIS/HCl buffer (pH 10.5) for 15 min at 4 °C. Absorbance was measured at 550 nm in FlexStation 3 microplate reader (Molecular Devices) or BioTek Synergy Neo2 microplate reader (Agilent).
LbuCas13a in vitro guide verification
The activity of LbuCas13a (GenScript) was monitored by measuring the fluorescence intensity of cleaved RNase Alert Reporter Substrates (Thermo Fisher Scientific). LbuCas13a: guide complex was formed by incubating 1 µM LbuCas13a with 0.5 µM gRNA in Cas13a reaction buffer (GenScript) at 37 °C for 10 min. After the complex was formed, several reaction conditions were prepared in a 384 black bottom plate. The final reaction concentrations for each component were 100 nM LbuCas13a:50 nM guide complex, 50 nM Target RNA, 180 nM RNase Alert Substrate, 1× Cas13a reaction buffer and 15 nM of RNase A. The reaction was incubated for 45 min at 37 °C, followed by measuring the fluorescence intensity (excitation λex. = 485 nm and λem. = 535 nm) using a BioTek Synergy H4 Hybrid plate reader. The background fluorescence was determined to be the reaction condition containing Cas13a buffer and RNase Alert Reporter, and was subtracted from the other reaction conditions.
RNA-seq analysis of transcript abundance
For the HeLa-GFP and HEK293-GFP cell lines, RNA sequencing and the fragments per kilobase of exon per million fragments mapped (FPKM) calculations were performed as described in a past work64. In brief, RNA isolation was performed using the NucleoSpin RNA Kit (Macherey-Nagel). The TruSeq RNA Library Prep Kit (v.2) from Illumina was used for library preparation, with an initial input amount of 500 ng of total RNA. The prepared libraries were sequenced with a 2 × 150 bp read length using the HiSeq 3000/4000 SBS Kit and an Illumina Hiseq 4000 sequencer. The adaptor trimmed, demultiplexed and quality filtered reads were aligned to the hg38 reference genome and transcriptome using Hisat2 (v.2.2.1). The Hisat2 output files (SAM) were converted to the BAM format and were sorted and indexed using SAMtools (v.1.3.1). The sorted BAM files were further processed using Cufflinks (v.2.1.1) to quantify the transcript abundances displayed in FPKM. For U2OS and NCI-H23 cell lines, RNA isolation was performed using the Quick-RNA Miniprep Kit (Zymo, R1054) according to the manufacturer’s protocol. Read-pair libraries were created with the NEBNext Ultra II Directional RNA Library Prep with rRNA Depletion kit (New England Biolabs, E7760). Next-generation sequencing of samples was subsequently performed on the Illumina NovaSeq X Series 150 × 150 bp platform. Sequencing reads were aligned to the GRCh38 genome (hg38) using the STAR aligner (v2.7.11b) with a pre-built RSEM index65. The RSEM reference index was generated using gene annotations from Gencode v.33 and the GRCh38 genome. Transcript abundance was quantified by RSEM, reporting FPKM values (v1.2.28)66. The STAR-aligned genome BAM output was sorted and indexed using SAMtools (v1.16)67.
LNP production and RNA delivery
LNPs were manufactured using Cytiva’s Ignite+ system equipped with NxGen mixer to provide 0.2 mg of formulated payload. The formulations were downstream processed and concentrations were adjusted to 0.25 mg ml–1. The mRNA and gRNA payload were co-packaged in a mass ratio of 60:40% (w/w). For the LNP application, HEK293T cells were seeded in 96-well plates. After 24 h, 40 ng of mRNA/gRNA-co-packaged LNPs were added to each 96-well containing ApoE3 (1 µg ml–1 final concentration, PeproTech) supplemented culture medium (DMEM containing 10% FCS and 4 mM Glutamine). After 6 h, the culture medium was replaced and the application of 40 ng mRNA/gRNA co-packaged LNPs was repeated. Confluence was quantified by Incucyte 72 h later.
Genomic DNA collateral cleavage assays
Collateral cleavage assays were performed as 10-μl reactions containing 0.1× NEB 3.1 buffer, 50 nM gRNA, 50 nM GeCas12a2, and nuclease-free water. After an initial 15-min incubation to form the RNP complex, 50 nM of target RNA were added to activate the complex. Approximately 50 ng of extracted genomic DNA (DNeasy Blood & Tissue Kit, Qiagen) was then added to each condition and samples incubated at 37 °C for 30 min. After 30 min, samples were treated with RNase A for 5 min and then Proteinase K for 10 min at 56 °C. Samples were visualized on a 1% TAE agarose gel run at 100 V for 40 min.
Immunofluorescence microscopy
GeCas12a2 or SuCas12a2 RNPs were delivered to 1 × 105 HeLa-GFP cells by electroporation which were then seeded onto glass coverslips and incubated for 18–24 h. Following incubation, samples were fixed with 4% paraformaldehyde in 1× PBS for 10 min then quenched with 100 mM glycine in 1× PBS for 10 min. Samples were rinsed with 1× PBS and subsequently permeabilized with 0.05% Triton-X in 1× PBS for 10 min. After an additional rinse with 1× PBS, samples were blocked with 2% BSA in 1× PBS for 1 h at room temperature. Following the block step and without rinsing, samples were treated with rabbit anti-53BP1 antibody (Novus Biologicals, NB100-306) diluted 1:1,000 in 1× PBS for 1 h at room temperature. Samples were then rinsed three times with 1× PBS before being treated with goat anti-rabbit Alexa Fluor 647 secondary antibody (Invitrogen, A32733) diluted 1:2,000 in 1× PBS for 1 h at room temperature. At the conclusion of the secondary incubation, samples were rinsed three times with 1× PBS waiting 5 min in between each rinse. Coverslips were mounted on glass slides using Prolong Diamond Antifade Mountant with DAPI (Thermo Fisher Scientific) overnight. Prepared samples were imaged using the Nikon Ti2 Eclipse confocal microscopy system with excitation of DAPI and 53BP1 fluorophore using 405 nm and 647 nm lasers, respectively. Raw images were exported from Nikon NIS-Elements software (v.5.41.02), split into individual channels using Fiji ImageJ software, and saved as .tif files. DAPI and 53BP1 channel images were then loaded into a custom CellProfiler pipeline to quantify the number of 53BP1 foci per cell.
Cell cycle analysis
Forty-eight hours after electroporation of GeCas12a2 or SuCas12a2 RNPs to 5 × 105 HeLa-GFP cells, cultures were suspended in 200 µl phenol red-free Leibovitz’s L−15 medium followed by 200 µl of 4% PFA fixative and incubated on ice for 25 min. To remove fixative, cells were pelleted by centrifugation at 250 RCF for 5 min and washed with 500 µl L-15 twice. To stain genomic DNA, cells were pelleted again and resuspended in 500 µl of a 1× PBS solution with 10 µg ml–1 DAPI stain (BD Pharmingen, 564907) and 0.1% Triton-X for 15 min at room temperature followed by 10 min on ice. Cells were strained through a 35-µm mesh cap before analysis to ensure single cell suspension. Ten thousand cells per condition were analysed by flow cytometry using the BD FACSCelesta system with 405 nm laser excitation for DAPI. Data were analysed using FlowJo v.10.
Annexin-V and caspase-3/7 staining
Forty-eight hours after electroporation of GeCas12a2 or SuCas12a2 RNPs to 5 × 105 HeLa-GFP cells, cultures were washed with 1× PBS, treated with 0.05% trypsin, centrifuged, and resuspended in annexin-V binding buffer (10 mM HEPES, 140 mM NaCl, 2.5 mM CaCl2, pH 7.4). For annexin V analysis, resuspended cells were mixed with Alexa Fluor 647 annexin-V antibody (BD Pharmingen, 567356) at a volumetric ratio of 1:40 and incubated at room temperature for 15–30 min. Immediately preceding data acquisition, samples were mixed with 10 µg ml–1 DAPI stain (BD Pharmingen, 564907) at a volumetric ratio of 1:80. For caspase-3/7 analysis, resuspended cells were mixed with 0.2 mM NucView 405 caspase-3/7 substrate (Biotium, 10407) at a volumetric ratio of 1:200 and incubated for 30 min at room temperature. Ten thousand cells per condition were analysed by flow cytometry using the BD FACSCelesta system with 405 nm laser excitation for DAPI or caspase-3/7 substrate and 640 nm laser excitation for annexin V. Data were analysed using FlowJo (v.10).
RNA-seq cell death pathway analysis
Forty-eight hours after electroporation of GeCas12a2 RNPs to 5 × 105 HeLa-GFP cells, total RNA was collected using Quick-RNA Miniprep Kit (Zymo, R1054) according to the manufacturer’s protocol. Read-pair libraries were created with NEBNext Ultra II Directional RNA Library Prep with poly(A) mRNA isolation kit (New England Biolabs, E7760). Next-generation sequencing of samples was then performed on the Illumina NovaSeq X Series 150 × 150 bp platform with 25 million reads per condition. Sequencing reads were aligned to hg38 with Bowtie2 (v.2.2.9)68. Read counts for each GENCODE gene were quantified with RSEM (v.1.2.28)66. We used DESeq2 (v.1.44.0)69 to normalize the read counts and perform differential expression analysis. We used the cut-off of padj <0.05 and fold change of >1.5 to identify genes significantly up- or downregulated between conditions indicated in the Article. Gene set enrichment analysis (v.4.3.3)70 was run to determine the enriched gene sets. We used the R package fgsea (v.1.30.0)71 to calculate the normalized enrichment statistics for each gene set in the Hallmark Collection from the Molecular Signatures Database72.
qPCR of incorporated dsODN for off-target double-strand-break detection
Detection of dsDNA breaks by qPCR of integrated donor oligos (dsODN) is adapted from the GUIDE-Seq method as described previously73. Amplicons were not sequenced in line with the full GUIDE-seq protocol, given expectations that the distribution of identified integration sites with the non-targeting guide would be difficult to distinguish from that of the vehicle-only control. One million cells were electroporated with 250 pmol of GeCas12a2 RNPs (or buffer) and 5 pmol of dsODN. The blunt-ended dsODN was prepared by annealing two single-stranded DNA oligos (dsODN_For/dsODN_Rev, Table S3). Forty-eight hours following electroporation, test conditions were collected for genome extraction using DNeasy Blood and Tissue Kit (Qiagen) according to the manufacturer’s protocol. Four hundred ng of collected genomic DNA were digested into 300–700 bp fragments and end-repaired using NEBNext Ultra II FS DNA Module (NEB) according to manufacturer’s protocol (10-min incubation at 37 °C). Reactions were purified using AMPure XP beads (Beckman Coulter) and ligated to adaptors (MNase_For/MNase_Rev, Table S3) with T4 Ligase (NEB). Reactions were purified using AMPure XP beads and PCR amplified with Illumina-based adaptors (PE_i5/PE_i7, Supplementary Table 3) using Q5 Master Mix (NEB). PCR-amplified samples were purified with AMPure XP beads before proceeding to qPCR on QuantStudio 3 (ThermoFisher) using Luna qPCR Master Mix (NEB) according to manufacturer’s protocol. Total input DNA Ct was determined by qPCR of each sample diluted 1:250 with primers against the Illumina-based adaptors (P5/P7, Supplementary Table 3). Amplification of dsODN-specific DNA was determined by qPCR of each sample with primers against one Illumina-based adaptor and the known dsODN sequence (PdsODN_For/PdsODN_Rev, Supplementary Table 3). Resulting Ct values were normalized to the Ct of respective total input DNA. Relative dsODN integration due to GeCas12a2 collateral activity was reported as the fold change from vehicle, which served as a baseline for background dsODN integration.
Cas12a2 off-target activation analysis
To investigate potential alternative targets of Cas12a2, the gRNA sequence under investigation was pasted into the Cas-OFFinder online tool74. The PAM SpRY Cas9 from Streptococcus pyogenes: 5′-NNN-3′ was selected and the genome searched was ‘Homo sapiens (GRCh38hg38)-Human’. As Cas12a2 targets RNA instead of DNA, the identified lower-mismatch-containing sequences were analysed using NCBI BLAST. This allowed us to identify the nucleotides surrounding the potential targeting region (including the 3′ PFS sequences), the location in the genome, and whether or not the sequence is part of an intrinsic region of DNA or an RNA transcript. To predict potential activating RNAs within the transcriptome, a cDNA search was performed using NCBI Nucleotide BLAST. The default settings were adjusted to increase the number of hits for short queries by selecting the minimum exact hit number (wordsize = 7), optimize for somewhat similar sequences and automatically adjust parameters for short input sequences. The Reference RNA sequences (refseq_rna) database was searched, filtering for the organism Homo sapiens (taxid:9606). The default settings were further adjusted by increasing the max target sequences to 5,000, with an expected threshold of 100, a match/mismatch score of 1,-3 and no filter or masking. To identify sequences with the most similarity to the target sequence, the MSA viewer was used to sort the hits by coverage percentage (smallest number of mismatches). To prevent any bias, no PFS restrictions were used. Instead, all potential activating RNAs were identified and reported (Supplementary Table 6).
Off-target in vitro testing
To determine the off-targets’ ability to activate the DNase activity of Cas12a2, a collateral dsDNA cleavage assay was performed, as previously described in this Article. However, as magnesium concentrations have been shown to affect Cas nucleases’ specificity, we lowered the magnesium concentration of the NEB 3.1 buffer to a more physiologically relevant magnesium concentration of 1 mM Mg2+ (100 mM NaCl, 50 mM Tris-HCl, 1 mM MgCl2, 100 µg ml–1 BSA, pH 7.9)75.
HeLa-GFP/-RFP co-culture cell depletion
GeCas12a2 or SuCas12a2 RNPs were electroporated to 1 × 105 cells containing HeLa-RFP cells and an excess of HeLa-GFP cells. After four days of culture incubation in 12-well plates, samples were resuspended and analysed by flow cytometry using the Sony MA900 system with 488 nm and 561 nm laser excitation. Quantification of relative GFP+/RFP+ cell populations excluded non-fluorescing cells as determined by gating strategy. Data were analysed using FlowJo (v.10).
Animal experiments
Fragments of patient-derived HPV16-positive HNSCC cancer xenograft specimens (HN11303) were thawed at 37 °C and transplanted subcutaneously into immunodeficient 5–7-week-old female NOG mice weighing 20.2 g ± 1.67 g. When tumours reached a volume of 1–1.5 cm3, the model was passaged into new mice. On the day of inoculation, tissue of passage 7 was collected. For transplantation of HN11303 fragments, tissue of passage 7 was collected and sliced into 3 × 3 to 4 × 4 mm3 pieces. These pieces were transplanted into the left flank of the anaesthetized experimental animals into a small pocket formed with scissors. From the first day onwards, tumour volumes were recorded two times per week. Animal welfare was controlled twice daily. In the treatment study, animals were randomized (n = 5 mice per group) and treatments were applied by intratumoral injections when an average tumour volume of about 150 mm3 was reached. The mice were treated on days 1 and 7, with two administrations per day spaced 6 h apart. In each treatment, 25 µl of the LNP formulation (containing 10 µg total RNA) or 25 µl of saline solution (vehicle control) was administered directly into the tumour. In the histology study, the animals were treated with one cycle (two injections with 6 h interval) only. Tumour volumes and mice body weights were measured twice per week. Tumour volume was calculated using the formula: tumour volume = (width2 × length)/2. As toxicity parameters, body weight, clinical signs, and animal behaviour were recorded for all mice twice a week. Data were not collected in a blinded manner. Mice were held in individual ventilated cages under standardized and controlled environmental conditions. Mice were housed at 24 ± 2 °C and 50 ± 10% relative humidity under an artificial 12 h light–dark cycle (lights on at 06:00 am, lights off at 6:00 pm). The experiment was terminated when the tumour size exceeded 1.5 cm3 as an ethical endpoint. The work conducted in living mice—at the Experimental Pharmacology and Oncology—is in accordance with the German Animal Welfare Act, and the UK Coordinating Committee on Cancer Research, and all procedures were approved by local authorities (Landesamt für Gesundheit und Soziales, LaGeSo) under approval no. E0023/23.
Indel enrichment
To induce indels in the GFP transgene, 1 × 105 HEK293-GFP cells were electroporated with purified FLAG-tagged FnCas12a (WP_003040289) RNPs. After electroporation (48 h), cells were detached from the 24-well plate cavities and transferred to poly-D-lysin-coated 10-cm-diameter culture dishes with the addition of 1 µg ml–1 tetracycline. One week after electroporation, cells were detached and subjected to indel analysis. To deplete unedited cells, 1 × 105 cells were electroporated with 50 pmol purified FLAG-tagged GeCas12a2 RNPs. After electroporation (48 h), cells were detached from the 24-well plate cavities and further cultivated on poly-D-lysin-coated 10-cm-diameter culture dishes with a medium containing 1 µg ml–1 tetracycline. After 10–12 days, the indel analysis was performed. For the indel analysis, genomic DNA was isolated (Quick-DNA Miniprep Plus Kit, Zymo Research), and the target region spanning parts of the cytomegalovirus (CMV) promoter and the GFP transgene was amplified by PCR using the CMV forward primer (5′-CCATAGTAACGCCAATAGGG-3′) and the GFP reverse primer (5′-TGTCGGCCATGATATAGACG-3′). The resulting amplicons were subjected to Sanger sequencing followed by determination of Indel-frequency via TIDE-analysis (https://tide.nki.nl/).
Prime edit enrichment
HEK293-GFP cells were seeded in a 24-well plate 24 h before transfection. Transfections were performed while cells were approximately 80% confluent. The transfection solution consisted of 1.5 µl X-tremeGENE HP DNA transfection reagent with 375 ng of PEmax-P2A-hMLH1dn plasmid, 125 ng of pegRNA plasmid and 41.5 ng of nicking sgRNA plasmid, these last two being cloned to target the GAPDH locus. The transfections were performed according to the manufacturer’s protocol. Samples were moved to six-well plates two days following transfection. Three days later, samples were electroporated with GeCas12a2 RNPs against the GAPDH target. Sample genomes were collected with DNeasy Blood and Tissue Kit (Qiagen) following manufacturer’s protocol 11 days after transfection (six days after electroporation). GAPDH amplicons for sequencing were prepared using primers 5′-TGAGTGCTACATGGTGAGCC-3′ and 5′-TGCAAAGAAAGAGGGAGCGG-3′. Editing efficiency was evaluated using long-read PCR sequencing (GENEWIZ).
Generation of KRAS and Cas12a2-expressing stable cell lines
Coding sequences for WT KRAS, KRASG12C and SuCas12a2 flanked by NLSs were cloned into a pLenti vector plasmid backbone including a P2A-GFP reporter; 10 µg of each expression plasmid, 5 µg of pMDLg/pRRE, 2.5 µg of pRSV-Rev and 2.5 µg of pMD2.G viral packaging plasmids were transfected into Lenti-X 293 T cells using polyethylenimine to produce the virus. The virus was collected, concentrated using Amicon Ultra-15 Centrifugal Filter Units (Millipore, UFC910024) and then used for transduction. Concentrated virus was added to cells in media containing 10 µg ml–1 polybrene; plates were gently shaken every 30 min for 3 h before the medium was replaced after 24 h. Cells with high GFP expression were sorted using the Sony MA900 cell sorting system.
RNA-FISH probe creation and RNA-FISH quantification of KRAS mRNA
Custom DNA probes against KRAS endogenous and overexpression constructs were designed using the Biosearch Technologies Stellaris RNA FISH Probe Designer. Designed DNA oligonucleotides were labelled with ddUTP-Cy5 (1 mM in water, Jena Bioscience, NU-1619-Cy5) using a Terminal Transferase Kit (NEB, M0315S). Labelled probes were purified and the labelling efficiency was measured using ultraviolet–visible absorbance. Cells were plated on glass coverslips in twelve-well plates and incubated for 24 h under standard culture conditions. Cells were washed with 1× PBS and fixed using 4% paraformaldehyde. Fixation was quenched with 0.1 M glycine in 1× PBS and permeabilized with 0.5% Triton X-100. Cells were rinsed again with 1× PBS and denatured using 10% formamide in 2× saline-sodium citrate (SSC) buffer. The coverslips were then placed on slides containing hybridization buffer containing 200 nM of the KRAS probe and incubated in a dark humidified chamber at 37 °C for 3 h. Following this incubation, coverslips were transferred to a new plate and incubated with 10% formamide in 2× SSC, once at 37 °C and once at room temperature. They were rinsed a final time with 1× PBS and then mounted on a slide cell side down using ProLong Diamond Antifade Mountant with DAPI (Invitrogen). Samples were incubated at room temperature for 18–24 h, sealed and then stored at 4 °C protected from light. To quantify the amount of KRAS transcripts in the cells, samples were imaged using a Nikon Eclipse Ti-2 microscope alongside a CSU-W1 spinning disk confocal scanner unit (Yokogawa). The samples were excited using four laser wavelengths: 405 nm, 488 nm, 561 nm and 640 nm. The fluorescence signals were captured using a Plan Apo 60× objective (Nikon, NA 1.40) and detected using a high-speed Kinetix sCMOS camera (photometrics). All imaging data were examined, processed and quantified using Fiji, Ilastik and Cell Profiler.
Generation of sotorasib-resistant cells
Sotorasib was purchased from MedChemExpress (Monmouth Junction). To achieve cellular resistance to this molecule, three biological replicates of NCI-H23 cells were exposed to progressively higher concentrations of sotorasib for one week each (1 μM, 2.5 μM, 5 μM and finally 10 μM); the surviving cells were propagated similarly to past protocols48.
Statistics and reproducibility
All statistical analyses and significance levels are defined in the figure legends. Exact P values for all comparisons are compiled in the Source data. Refer to Supplementary Fig. 1 for gel source data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.